Introducción
Constructing a genomic DNA library is a foundational technique in molecular biology, essential for a wide range of genetic explorations. This process involves several critical steps:
- Isolating
- Fragmenting
- Ligating
- Transforming
- Screening
Mastering these steps allows researchers to gain vital insights into genetic structures and functions.
However, each step presents its own complexities and challenges. Researchers often encounter questions regarding best practices and potential pitfalls during the construction of genomic libraries. Understanding the key techniques that ensure successful library construction is crucial. By navigating these complexities effectively, researchers can enhance their genetic investigations and contribute to advancements in the field.
Isolate Genomic DNA
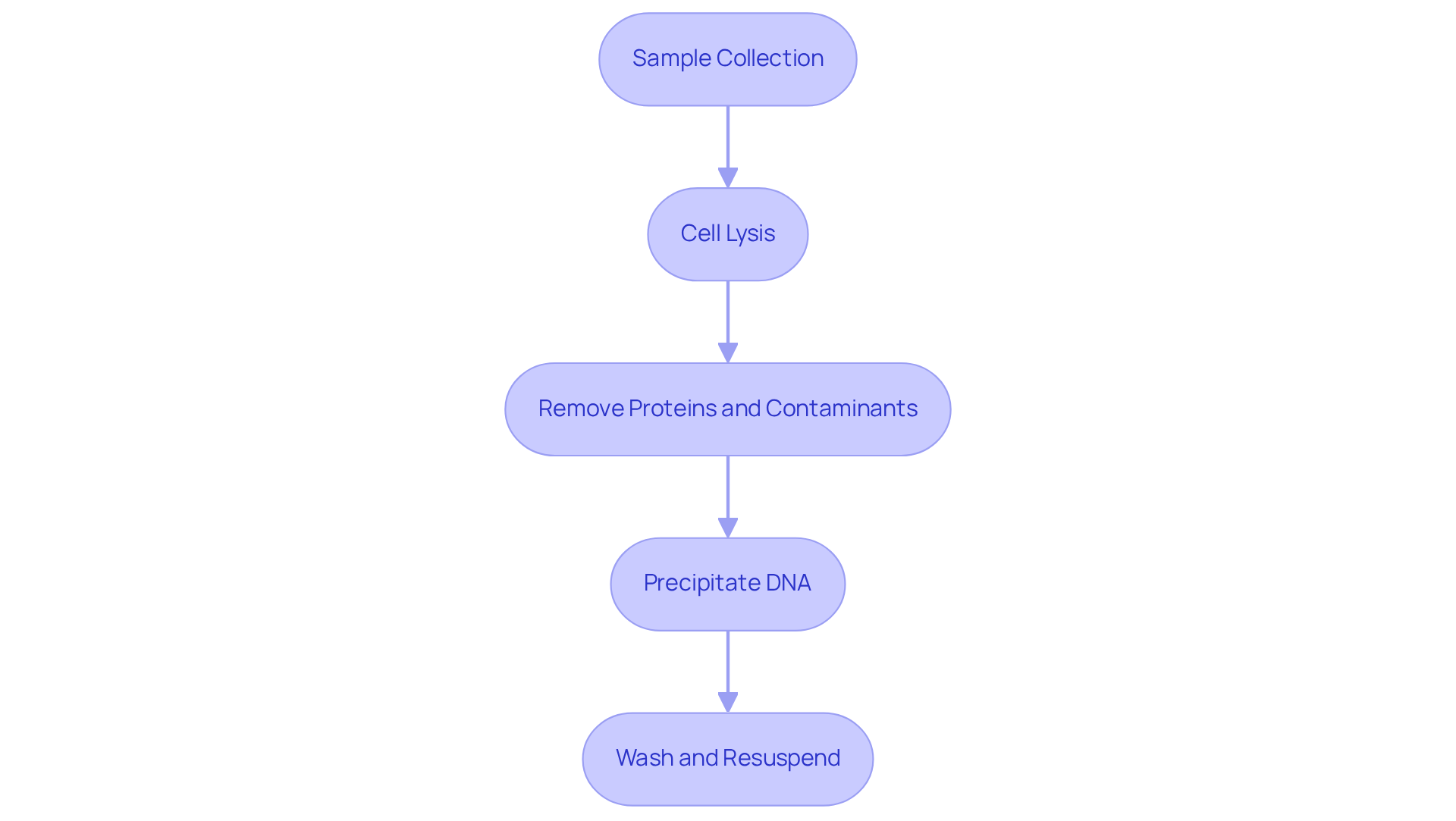
To isolate genomic DNA, follow these structured steps:
-
Sample Collection: Begin by gathering tissue or biological samples from the organism of interest. It is crucial to ensure that these samples are stored properly to prevent degradation.
-
Cell Lysis: Utilize a lysis buffer that contains detergents and enzymes, such as Proteinase K, to effectively break down cell membranes and release the DNA. This process typically involves incubating the sample at 56°C for a duration of 1-2 hours.
-
Remove Proteins and Contaminants: Introduce phenol-chloroform to the lysate to facilitate the separation of proteins from the DNA. Following this, centrifuge the mixture and carefully transfer the aqueous layer, which contains the DNA, to a new tube.
-
Precipitate DNA: To precipitate the DNA, add cold ethanol or isopropanol to the aqueous solution. It is advisable to incubate this mixture at -20°C for at least 30 minutes.
-
Wash and Resuspend: After centrifuging to collect the DNA pellet, wash it with 70% ethanol and allow it to air dry. Finally, resuspend the DNA in a suitable buffer, such as TE buffer, for optimal storage.
Fragment Genomic DNA
To effectively fragment genomic DNA, it is essential to follow these steps:
-
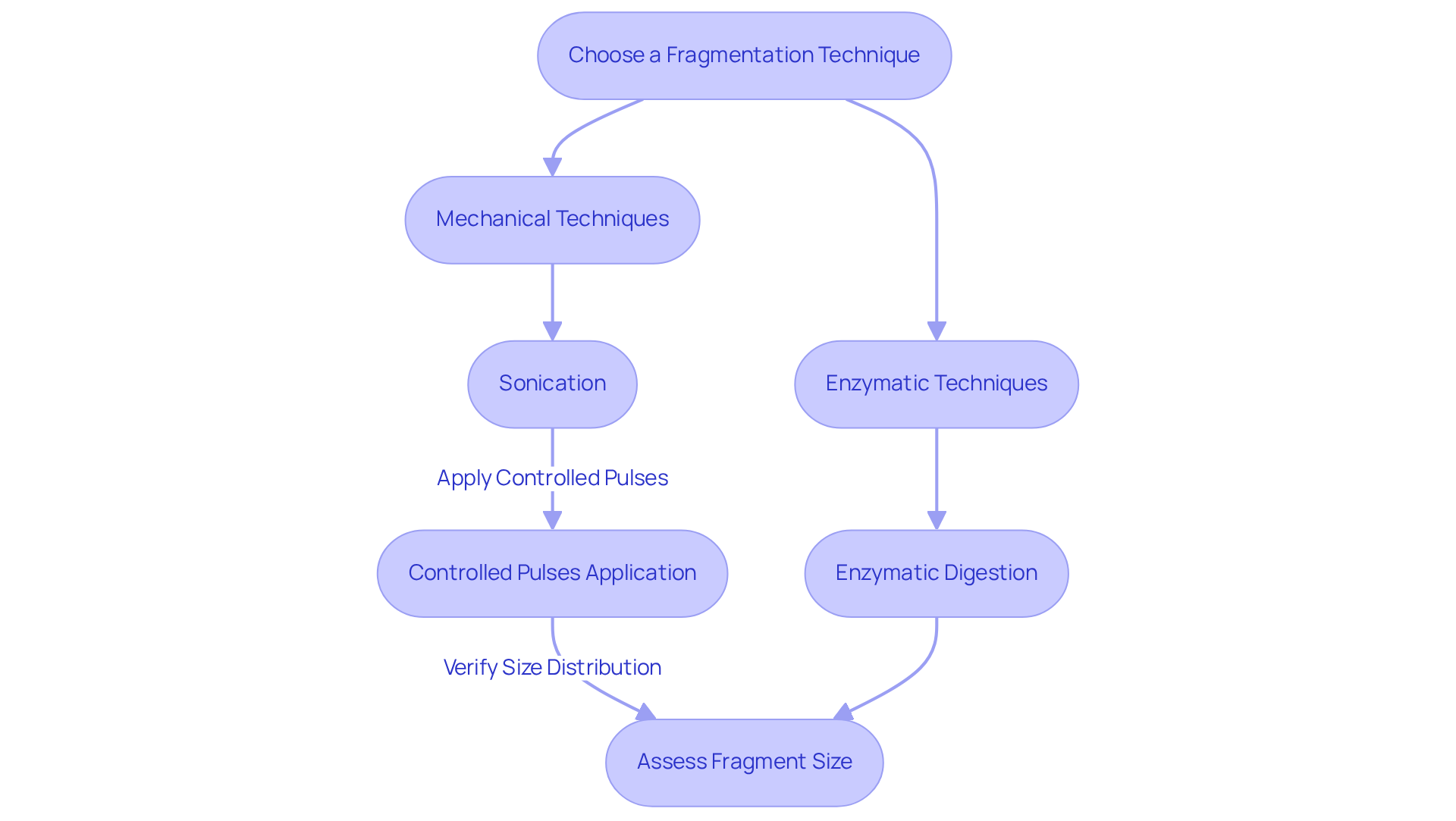
Choose a Fragmentation Technique: You can select between mechanical techniques, such as sonication, hydrodynamic shearing, or nebulization, which yield random fragmentation, or enzymatic approaches, like restriction enzymes, which can produce specific fragment sizes tailored to your needs.
-
Sonication: For sonication, place the DNA solution in a sonicator and apply controlled pulses to shear the DNA into smaller fragments. Aim for a typical fragment size of 200-500 bp, adjusting the duration and intensity of the pulses as necessary to achieve the desired outcome.
-
Enzymatic Digestion: When selecting enzymatic techniques, choose suitable restriction enzymes that target specific sequences within the DNA. Combine the DNA with the enzyme and a suitable buffer, then incubate at the enzyme’s optimal temperature to facilitate precise cutting.
-
Assess Fragment Size: To verify the size distribution of the fragmented DNA, run a small aliquot on an agarose gel. This step is crucial for confirming that the fragmentation conditions have produced the desired fragment sizes, allowing for adjustments if necessary.
Current trends indicate that mechanical fragmentation techniques, particularly sonication, hydrodynamic shearing, and nebulization, are favored for their ability to produce a wide range of fragment sizes and enhance library complexity. In contrast, enzymatic approaches are appreciated for their specificity and consistency, with studies demonstrating that they can achieve high yields even with low input quantities. For instance, NEBNext Ultra II FS DNA libraries can generate fragments with input as low as 100 pg, showcasing the efficiency of these techniques in modern genomic applications. Successful applications of both approaches in laboratories emphasize their respective benefits, making them essential for the construction of genomic DNA library. Furthermore, the concept of molecular randomness is crucial, as it indicates the absence of systematic cleavage bias, leading to a Poisson distribution of breakpoints, which is vital for reliable downstream analyses. The advancements in DNA fragmentation techniques have been pivotal since the Human Genome Project, underscoring the importance of these methods in genomic research.
Ligate DNA into Vector System
To ligate DNA into a vector, adhere to the following structured steps:
-
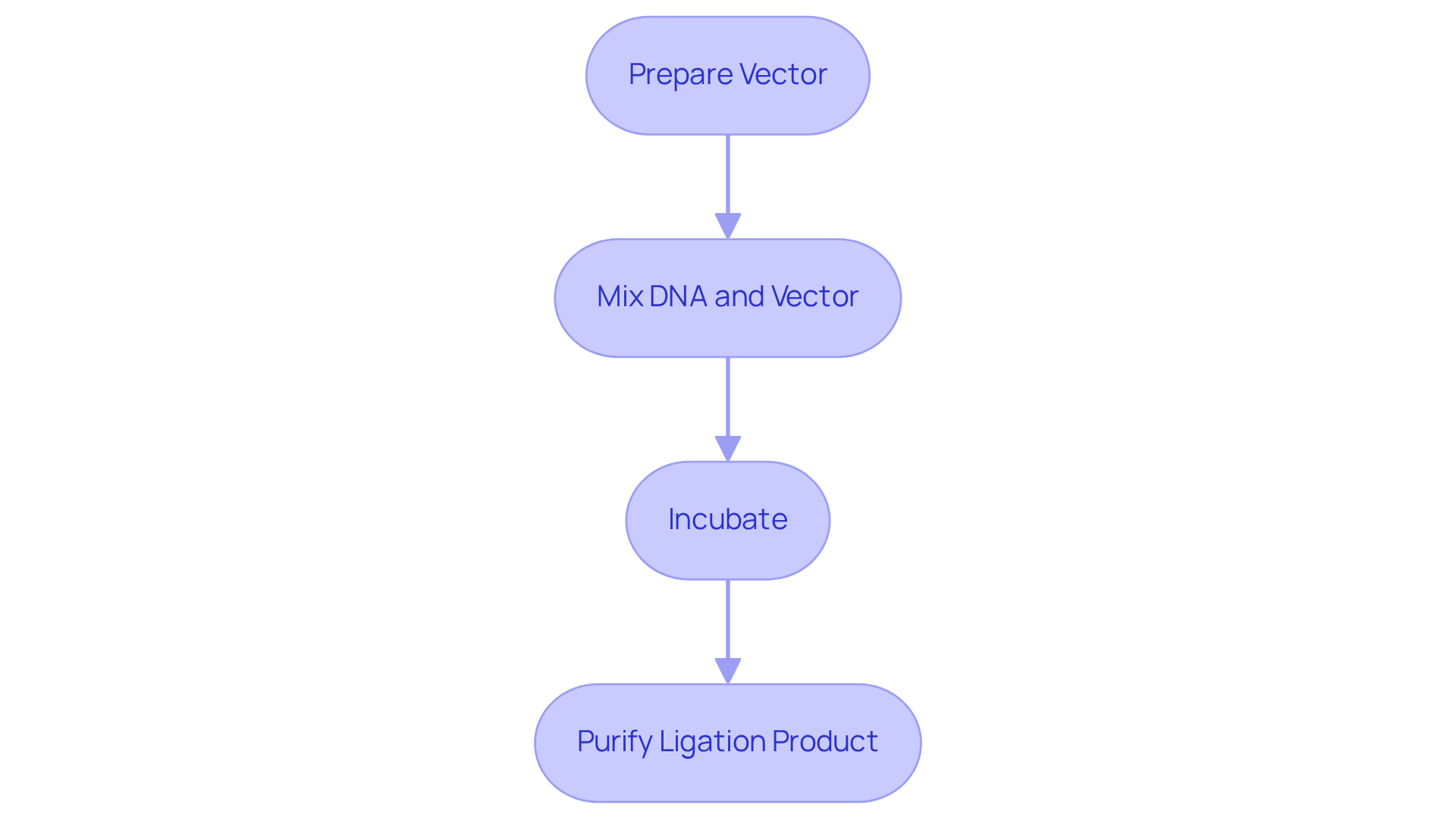
Prepare Vector: Begin by linearizing the vector through digestion with the same restriction enzyme utilized for fragmenting the genomic DNA. It is essential to purify the digested vector for the construction of genomic DNA library to eliminate enzymes and contaminants.
-
Mix DNA and Vector: In a microcentrifuge tube, combine the DNA fragments with the linearized vector in a molar ratio of 3:1 (insert:vector). Introduce T4 DNA ligase along with the appropriate buffer to facilitate the construction of genomic DNA library.
-
Incubate: Allow the mixture to incubate at room temperature for a duration of 10 to 30 minutes, or alternatively, at 16°C overnight to achieve optimal ligation efficiency.
-
Purify Ligation Product: Following the ligation, purify the resultant product using a PCR cleanup kit to remove any unligated DNA and enzymes. This purification step is crucial for ensuring the success of the subsequent transformation in the construction of a genomic DNA library.
Transform Recombinant DNA into Host Cells
To successfully transform recombinant DNA into host cells, follow these structured steps:
-
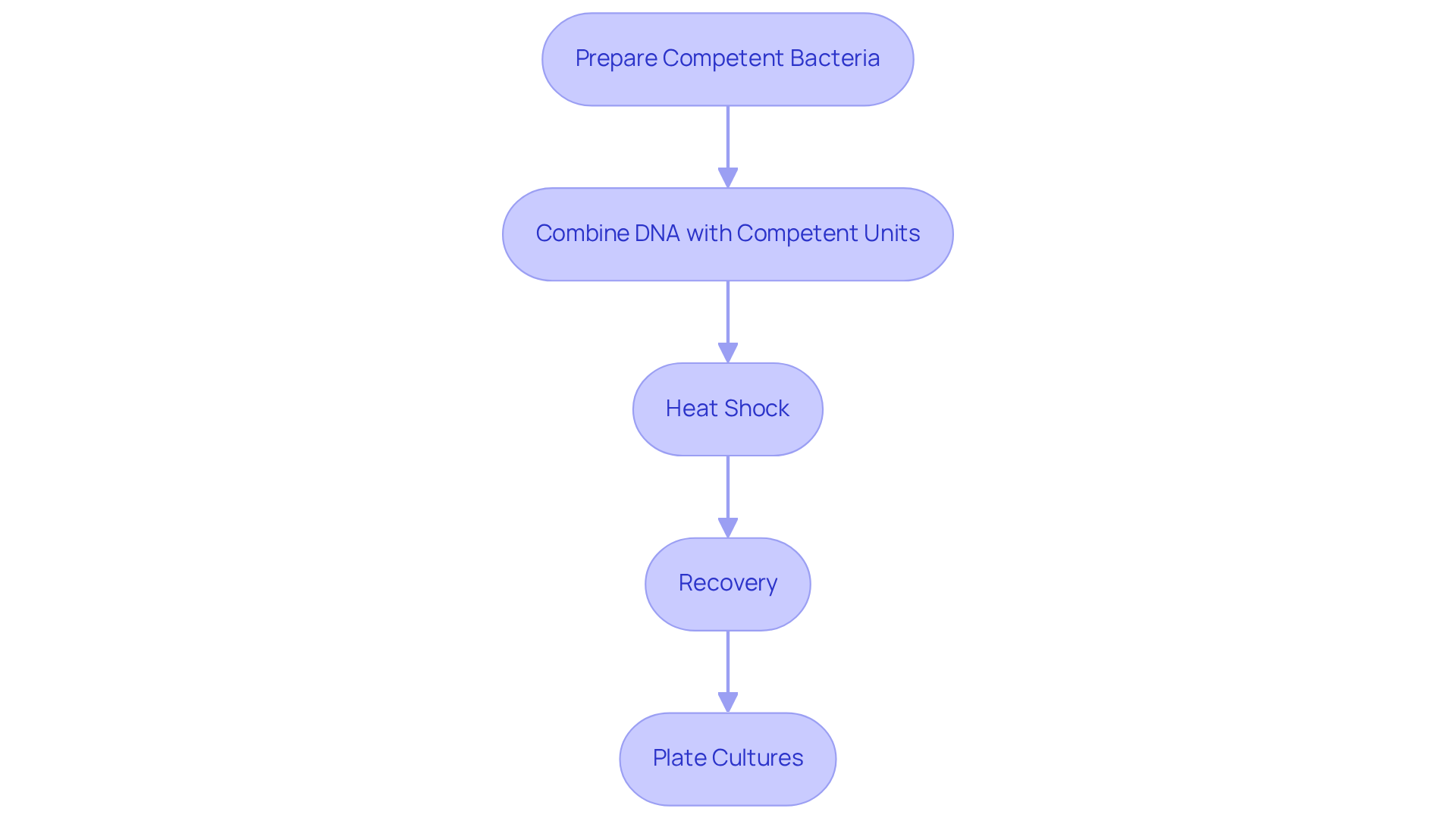
Prepare Competent Bacteria: Utilize commercially available chemically competent E. coli bacteria or create your own by treating the organisms with calcium chloride. This treatment enhances the permeability of the membrane, facilitating DNA uptake.
-
Combine DNA with Competent Units: Introduce 1-5 µL of the ligation product into 50 µL of competent units within a microcentrifuge tube. Gently mix the contents by flicking the tube to ensure even distribution of the DNA.
-
Heat Shock: Place the mixture on ice for 30 minutes, then subject it to a heat shock by immersing the tube in a 42°C water bath for 45-60 seconds. This critical phase encourages the absorption of DNA into the bacterial cells.
-
Recovery: Immediately return the tube to ice for 2 minutes, then add SOC medium and incubate at 37°C for 1 hour. This recovery period allows the organisms to express the antibiotic resistance gene effectively. Recent advancements in gene editing technologies, including the FDA’s approval of CRISPR-Cas9-based gene therapy, underscore the significance of this step in ensuring successful transformation.
-
Plate Cultures: Distribute the altered cultures onto agar plates containing the appropriate antibiotic and incubate overnight at 37°C to promote colony growth. Average recovery times for transformed E. coli cells typically range from 30 minutes to 1 hour, depending on the specific protocol employed.
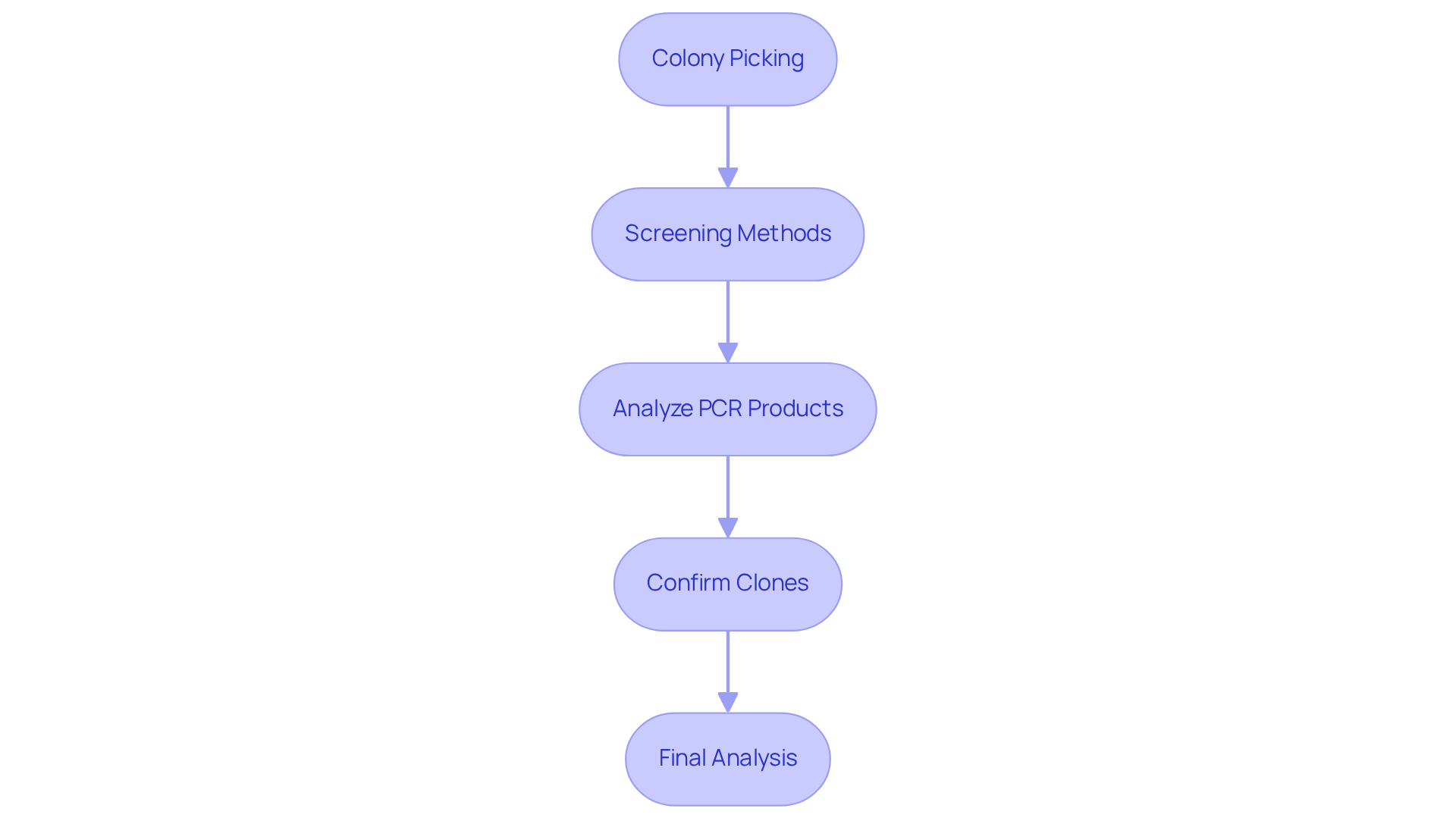
Screen Genomic Library
To effectively screen your genomic library, follow these structured steps:
-
Colony Picking: After incubation, select colonies from agar plates cultured with antibiotics. Each colony represents a potential clone that is essential for the construction of genomic DNA library containing recombinant DNA.
-
Screening Methods: Utilize techniques such as colony PCR, hybridization, or restriction analysis to identify the presence of the insert. Colony PCR is particularly efficient, as it uses a small sample from the colony as a template for amplification.
-
Analyze PCR Products: Subject the PCR products to agarose gel electrophoresis to verify the presence and size of the insert. Compare the results against a DNA ladder to confirm that the expected size is present. Be aware that polysaccharides and pigments can impair DNA purity, potentially affecting PCR success.
-
Confirm Clones: For clones that test positive, conduct additional analyses, such as sequencing, to validate the identity of the insert and ensure it aligns with the intended sequence.
Utilizing colony PCR has demonstrated high success rates, particularly in the construction of genomic DNA library, with studies indicating an overall PCR success of 86% across various fungal taxa. This approach not only streamlines the screening process but also enhances the reliability of identifying recombinant clones. However, it is crucial to note that optimized methods may be necessary for specific fungal groups to achieve consistent results.
Conclusión
Constructing a genomic DNA library is a foundational aspect of modern genetics and molecular biology, representing both a technical challenge and an opportunity for significant advancements. By adhering to systematic methods for isolating genomic DNA, fragmenting it, ligating it into vector systems, transforming it into host cells, and screening the resulting library, researchers can create a robust genomic resource for further analysis and experimentation.
This article details each phase of the process. It begins with the careful isolation of genomic DNA, ensuring quality and integrity. Next, it discusses fragmentation techniques that generate suitable DNA sizes for ligation. The importance of proper vector preparation and ligation conditions is emphasized, along with the necessary steps for transforming recombinant DNA into host cells. Finally, the article highlights the screening processes that confirm successful cloning, which is essential for validating the integrity of the genomic library.
In summary, by applying the steps outlined in this guide, researchers can enhance their capabilities in genomic studies. This structured approach not only paves the way for discoveries that could lead to advancements in biotechnology and medicine but also ensures that genomic libraries are constructed with precision. Ultimately, this contributes significantly to the broader field of genetic research and its applications.
Preguntas frecuentes
What are the steps to isolate genomic DNA?
To isolate genomic DNA, follow these steps: collect tissue or biological samples, use a lysis buffer to break down cell membranes, separate proteins from DNA using phenol-chloroform, precipitate the DNA with cold ethanol or isopropanol, and finally wash and resuspend the DNA in a suitable buffer.
What is the purpose of cell lysis in DNA isolation?
Cell lysis is performed to break down cell membranes and release the DNA contained within the cells, allowing for further processing and purification.
How do you remove proteins and contaminants from the DNA lysate?
Introduce phenol-chloroform to the lysate, centrifuge the mixture, and transfer the aqueous layer, which contains the DNA, to a new tube.
What is the process for precipitating DNA?
To precipitate DNA, add cold ethanol or isopropanol to the aqueous solution and incubate it at -20°C for at least 30 minutes.
How should the DNA be washed and stored after isolation?
After centrifuging to collect the DNA pellet, wash it with 70% ethanol, allow it to air dry, and then resuspend it in a suitable buffer, such as TE buffer, for optimal storage.
What techniques can be used to fragment genomic DNA?
Fragmentation techniques include mechanical methods like sonication, hydrodynamic shearing, or nebulization, and enzymatic approaches using restriction enzymes.
How does sonication work for DNA fragmentation?
In sonication, the DNA solution is placed in a sonicator, where controlled pulses are applied to shear the DNA into smaller fragments, targeting a typical size of 200-500 bp.
What is the role of restriction enzymes in enzymatic digestion?
Restriction enzymes are selected to target specific sequences within the DNA, allowing for precise cutting and generation of specific fragment sizes.
How can you assess the size of fragmented DNA?
To verify the size distribution of fragmented DNA, run a small aliquot on an agarose gel, which helps confirm that the fragmentation conditions produced the desired fragment sizes.
What are the current trends in DNA fragmentation techniques?
Current trends favor mechanical fragmentation techniques for their ability to produce a wide range of fragment sizes, while enzymatic approaches are valued for their specificity and high yields even with low input quantities.
Why is the concept of molecular randomness important in DNA fragmentation?
Molecular randomness indicates the absence of systematic cleavage bias, leading to a Poisson distribution of breakpoints, which is crucial for reliable downstream analyses in genomic research.





