Einführung
The complexity of genomic data analysis poses significant challenges that can affect patient outcomes. This guide will break down the process of genomic data analysis into six manageable steps, equipping you with the necessary skills and knowledge. A structured approach is essential to mitigate risks and ensure ethical compliance in genomic analyses.
Understand the Fundamentals of Genomic Data Analysis
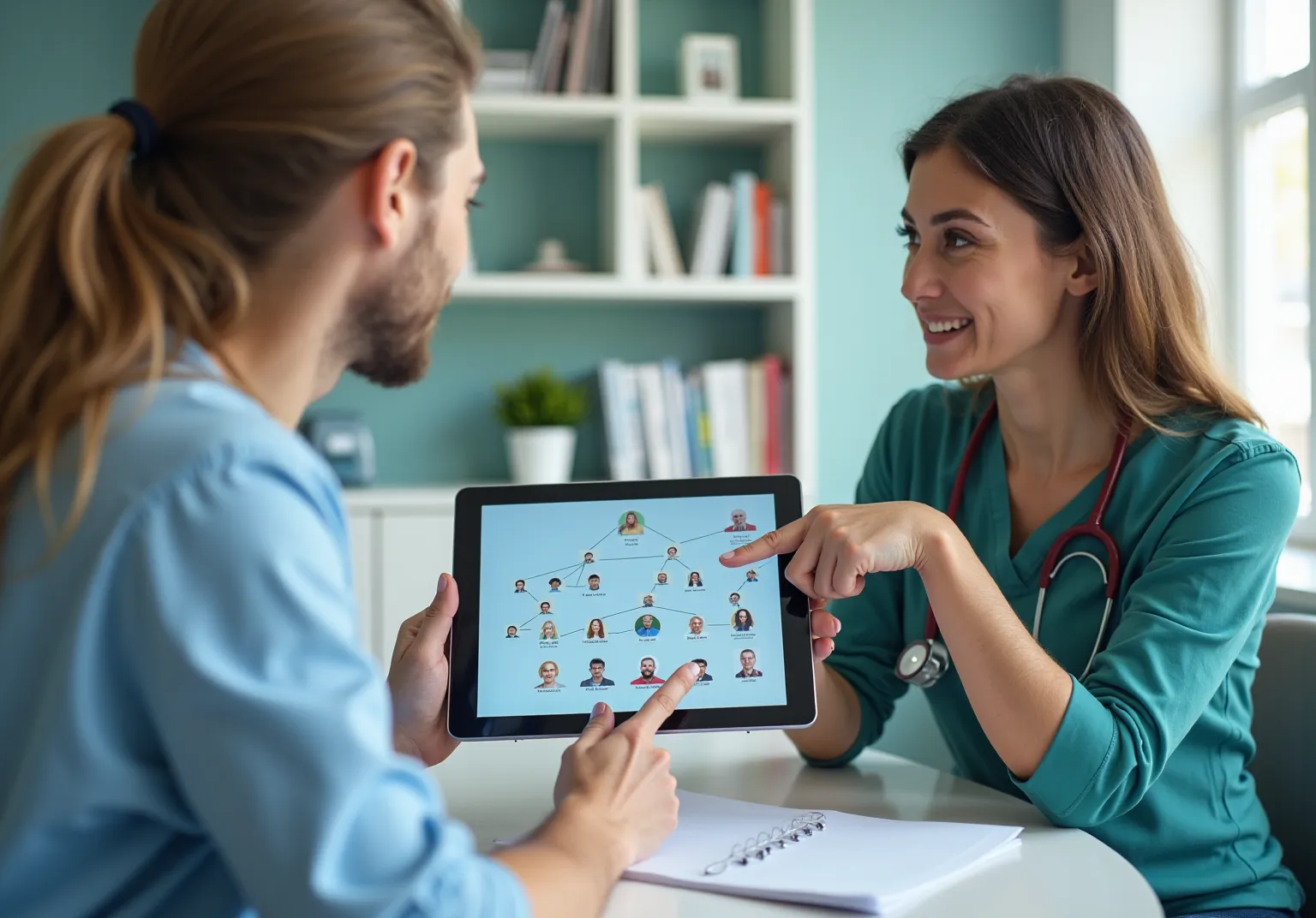
Understanding genomic data is crucial for effectively interpreting sequencing results and their implications for health. Key concepts include:
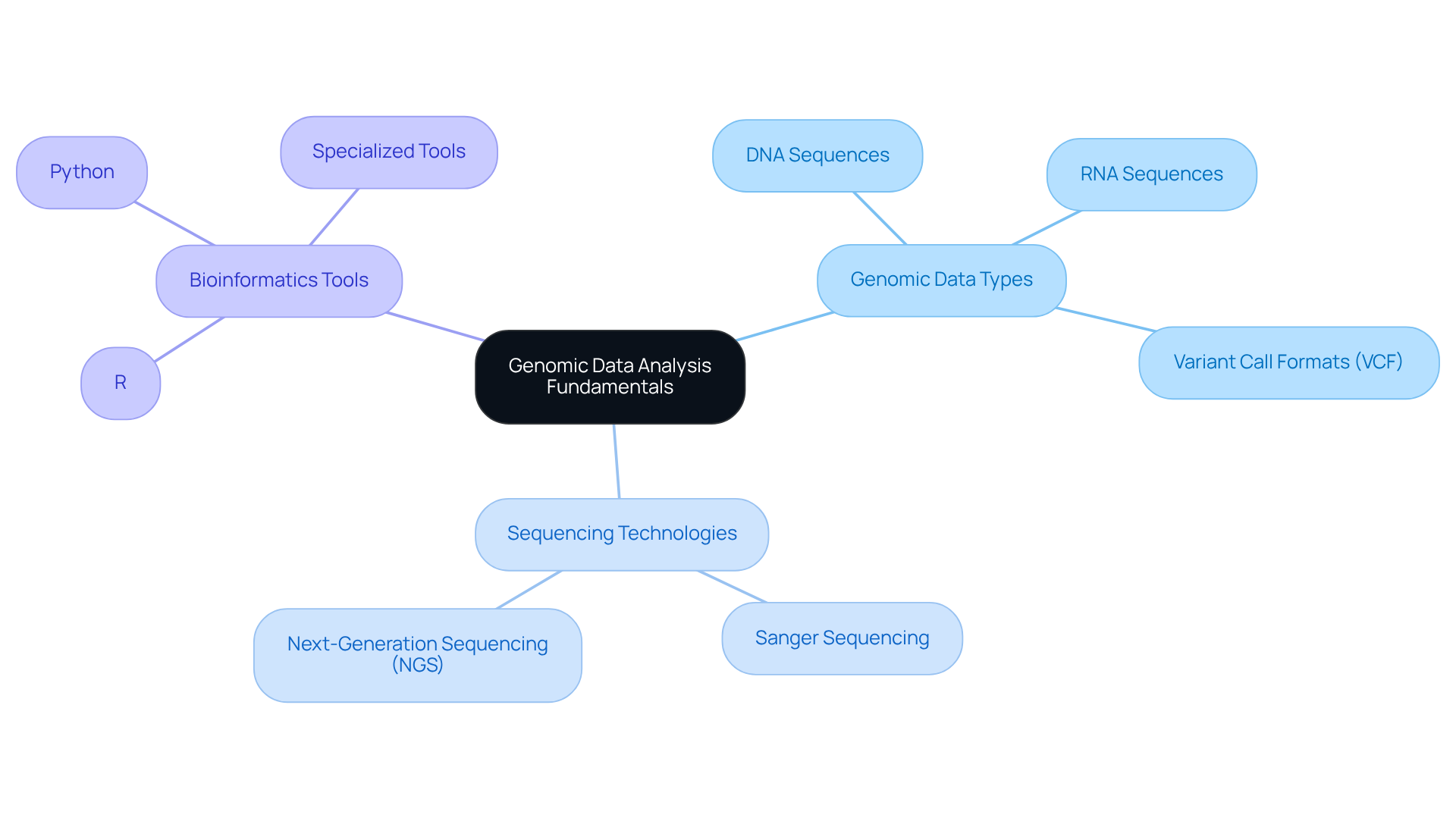
- Genomic Data Types: Familiarity with various genomic data types, including DNA sequences, RNA sequences, and variant call formats (VCF), is essential for effective evaluation and application in clinical settings.
- Sequencing Technologies: A comprehensive grasp of sequencing methods, such as Sanger sequencing and Next-Generation Sequencing (NGS), is vital. NGS, in particular, has transformed genetic studies by facilitating high-throughput sequencing, which is crucial for extensive research and the generation of genomic data in clinical diagnostics.
- Bioinformatics Tools: Proficiency in bioinformatics tools is essential for genetic study. Widely used software and coding languages, like R and Python, along with specialized bioinformatics tools, aid in the evaluation and interpretation of intricate genetic information.
By 2026, it is expected that 70% of healthcare workers will utilize bioinformatics tools in genetic studies, highlighting the growing role of technology in healthcare. TrakGene enhances this integration by providing comprehensive electronic health records related to genetics, including cancer risk tools, enabling a more streamlined approach to managing genetic information and improving patient experience. For instance, the collaboration between PacBio and Children’s Mercy Kansas City showcases how advanced sequencing technologies can deepen our understanding of genetic diseases and how TrakGene’s solutions can support similar progress. Mastering these fundamentals not only equips you to navigate the complexities of genetic information evaluation but also positions you to drive advancements in personalized medicine.
Collect Relevant Genomic Data
Effective genomic data analysis hinges on a systematic approach to data collection:
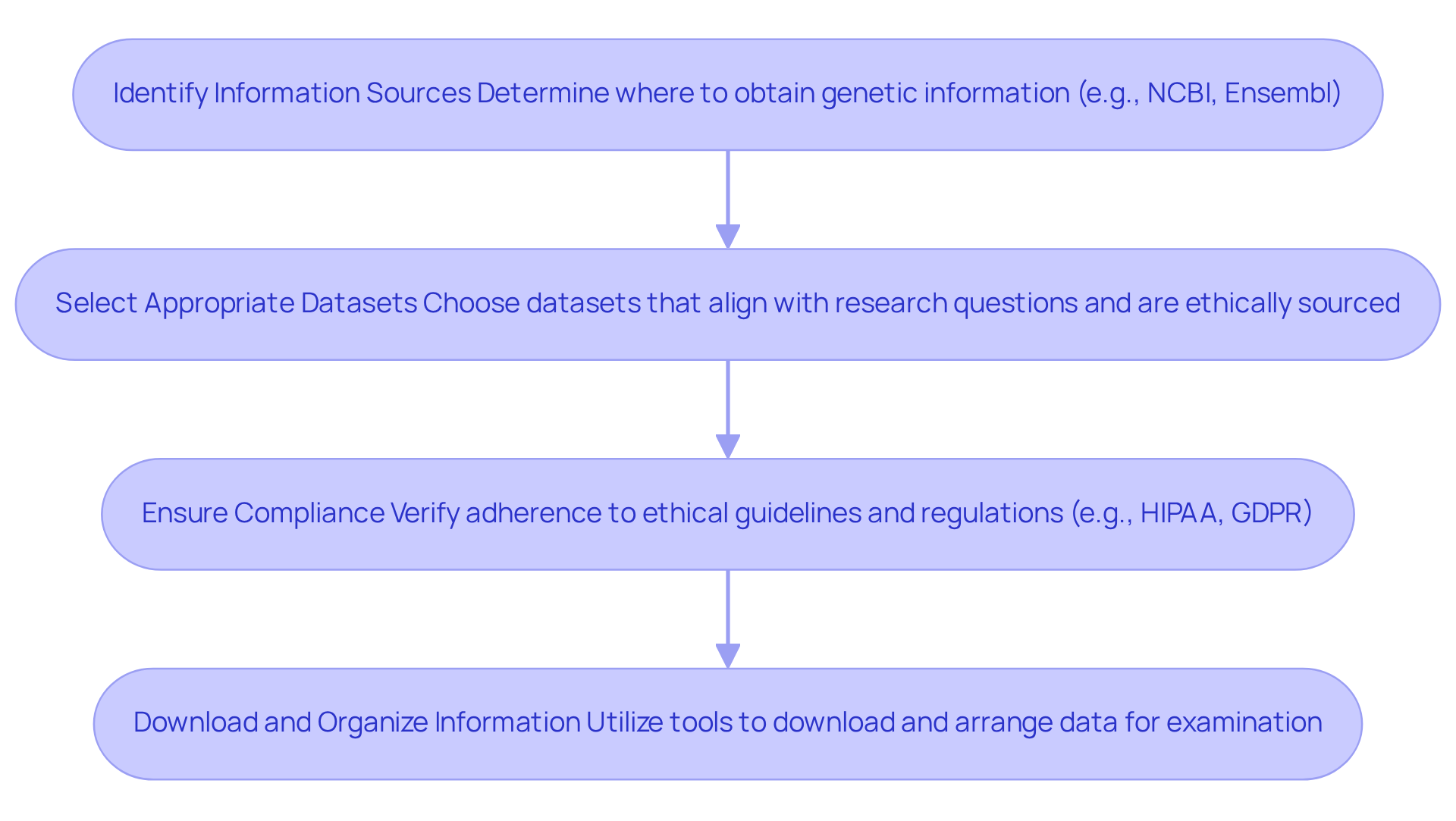
- Identify Information Sources: Determine where to obtain genetic information, such as public databases like NCBI and Ensembl, or institutional repositories that house valuable collections.
- Select Appropriate Datasets: Choose datasets that align with your research question or clinical needs, ensuring they are relevant, comprehensive, and ethically sourced.
- Ensure Compliance: Verify that information collection adheres to ethical guidelines and regulations, including informed consent and privacy laws such as HIPAA and GDPR. In 2026, compliance rates with these regulations are critical for maintaining trust and integrity in genomic research. Non-compliance can lead to significant repercussions, undermining the credibility of the research.
- Download and Organize Information: Utilize appropriate tools to download the information in formats suitable for examination (e.g., FASTQ, BAM). Arrange the information methodically to enable straightforward access during examination.
Adhering to these guidelines ensures the creation of a robust dataset, vital for accurate analysis of genomic data. The World Health Organization stresses the significance of informed consent and privacy as essential components in genetic information collection, ensuring that individuals comprehend and consent to the use of their genetic details. Ultimately, adherence to ethical standards not only safeguards individual rights but also enhances the credibility of genomic research.
Perform Data Quality Checks and Cleaning
Ensuring the integrity of genomic data is crucial; however, many overlook the necessary quality checks that can prevent significant errors. Follow these steps for effective quality checks and data cleaning:
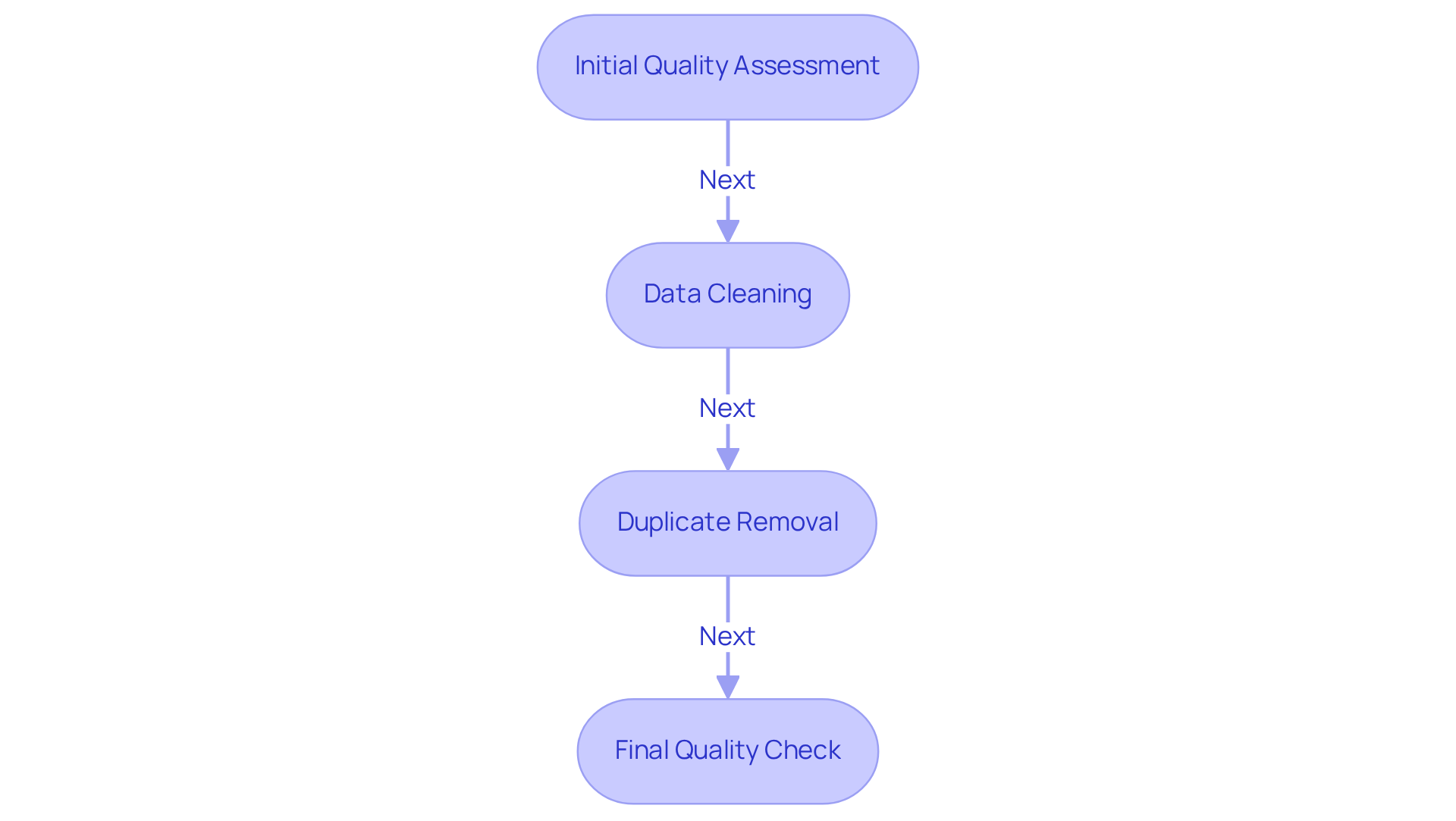
- Initial Quality Assessment: Utilize tools like FastQC to evaluate the quality of raw sequencing information, checking for issues such as low-quality reads, adapter contamination, and GC content.
- Data Cleaning: Implement cleaning techniques to remove low-quality reads and contaminants. Tools like Trimmomatic can help trim sequences based on quality scores.
- Duplicate Removal: Identify and eliminate duplicate reads to avoid bias in the evaluation. Tools like Picard can assist in this process.
- Final Quality Check: After cleaning, conduct a final quality evaluation to ensure that the information meets the necessary standards for examination.
Neglecting these quality checks could compromise the accuracy of your evaluations of genomic data, leading to potentially misleading conclusions.
Process Genomic Data for Analysis
Preparing genomic data for analysis requires a systematic approach to ensure accuracy and reliability:
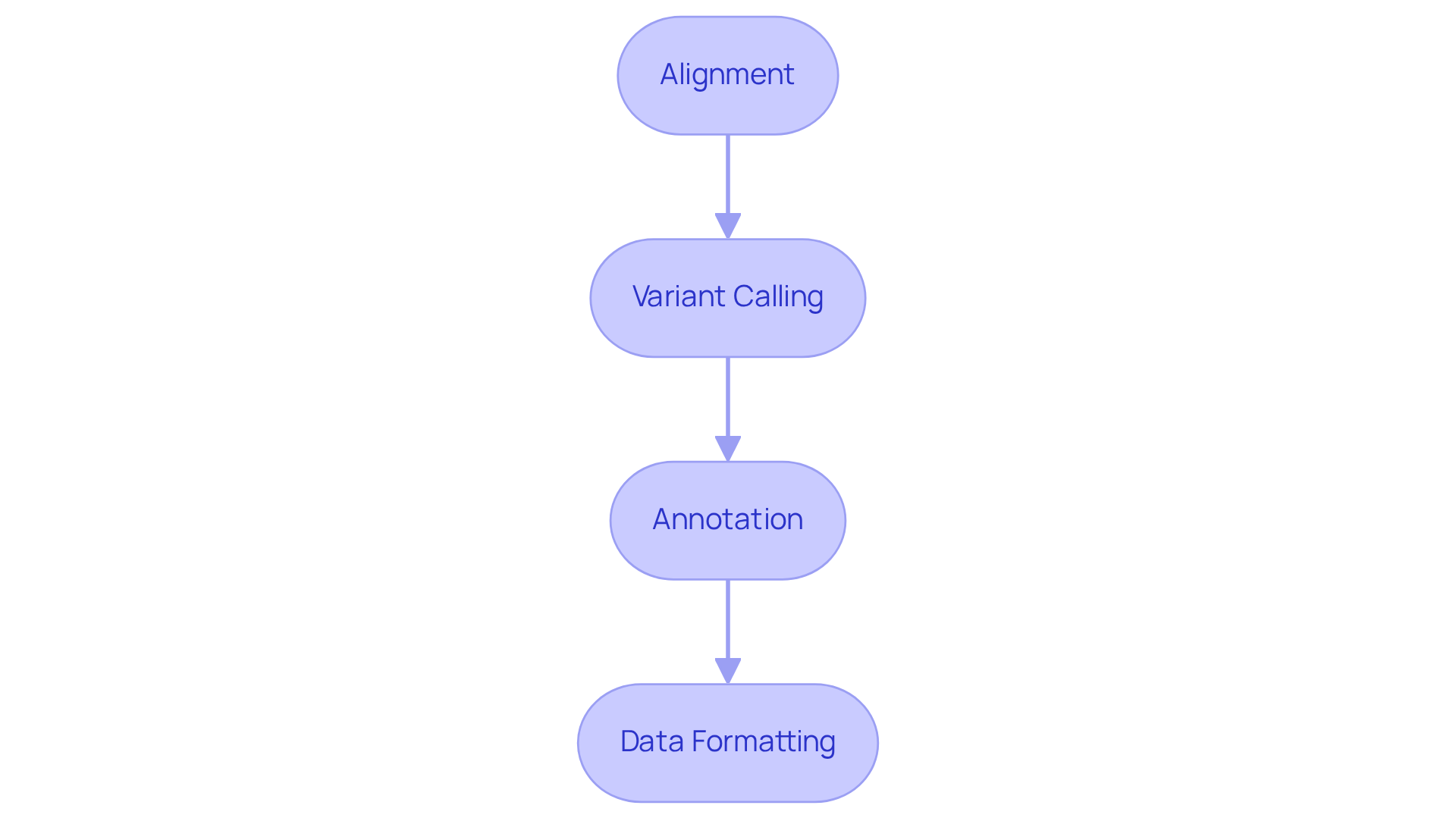
- Alignment: Utilize alignment tools such as BWA or Bowtie to align sequencing reads to a reference genome. This phase is crucial for identifying variants, as the alignment technique affects downstream analysis.
- Variant Calling: Employ variant calling software like GATK or FreeBayes to identify genetic variants from the aligned data. This process generates a Variant Call Format (VCF) file, which contains detailed information about the identified variants. Recent updates in variant calling software have enhanced accuracy, with studies showing that the Mendelian Inheritance Error Rate (MIER) varies by method.
- Annotation: Annotate the identified variants using established databases such as dbSNP or ClinVar. This step provides critical context regarding the biological significance of the variants, aiding in the interpretation of their potential impact on health.
- Data Formatting: Convert the processed information into formats that enable examination, such as transforming VCF files into tabular formats. This conversion improves information handling and integration with other analytical tools.
By following these steps, you ensure that your genomic data is primed for thorough analysis, which ultimately enhances the reliability of your findings.
Conduct Exploratory Data Analysis and Modeling
To achieve meaningful insights in genomic research, a structured approach to exploratory data analysis (EDA) and modeling is essential:
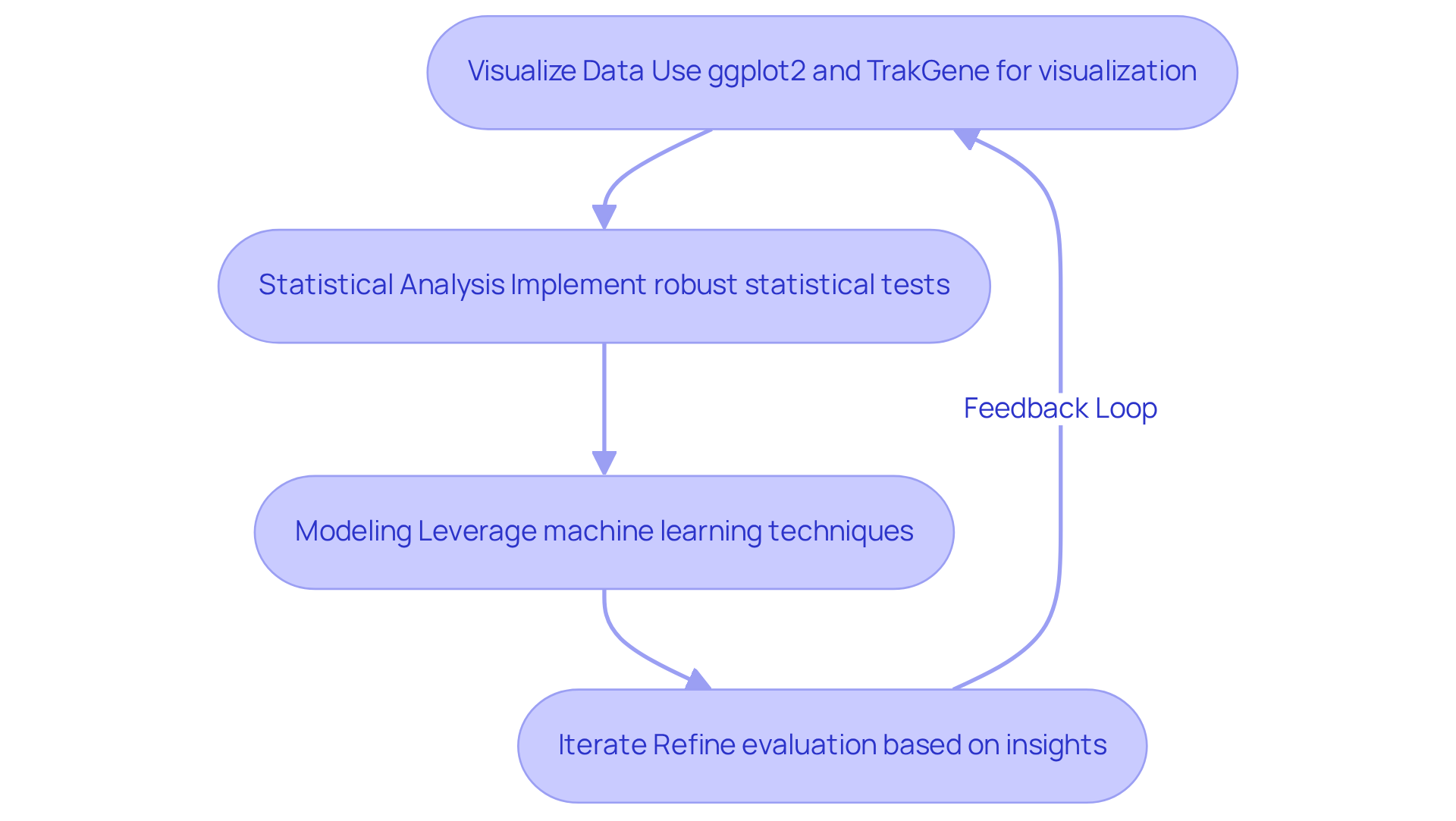
- Visualize Data: Utilize advanced visualization tools, such as ggplot2 in R and TrakGene’s Clinical Genetics Management Software, to create informative plots that reveal trends and patterns within your dataset. Effective visualization is essential for understanding complex genomic data and can be simplified through TrakGene’s intuitive interface.
- Statistical Analysis: Implement robust statistical tests to evaluate relationships between variables. Techniques such as correlation evaluation and t-tests can provide insights into the significance of observed patterns, aiding in hypothesis testing. TrakGene’s software can automate these analyses, enhancing efficiency and accuracy.
- Modeling: Leverage machine learning techniques, including regression and clustering, to develop predictive models. These models can reveal potential relationships and outcomes, enhancing the understanding of genomic data. Data scientists emphasize that ensemble methods can significantly boost model performance by integrating predictions from various algorithms, a feature supported by TrakGene’s software.
- Iterate: Continuously refine your evaluation based on insights gained from EDA. Adjust your models and visualizations as necessary to improve accuracy and relevance. Kasey Couts, PhD, notes that modern genetic assessment software like TrakGene allows researchers to focus on biological insights rather than getting bogged down by complex bioinformatics challenges.
Ultimately, a thorough EDA process not only enhances understanding of genomic data but also drives informed decisions in genomic research.
Visualize and Report Genomic Analysis Findings
To ensure effective communication of genomic analysis findings, a structured approach is essential:
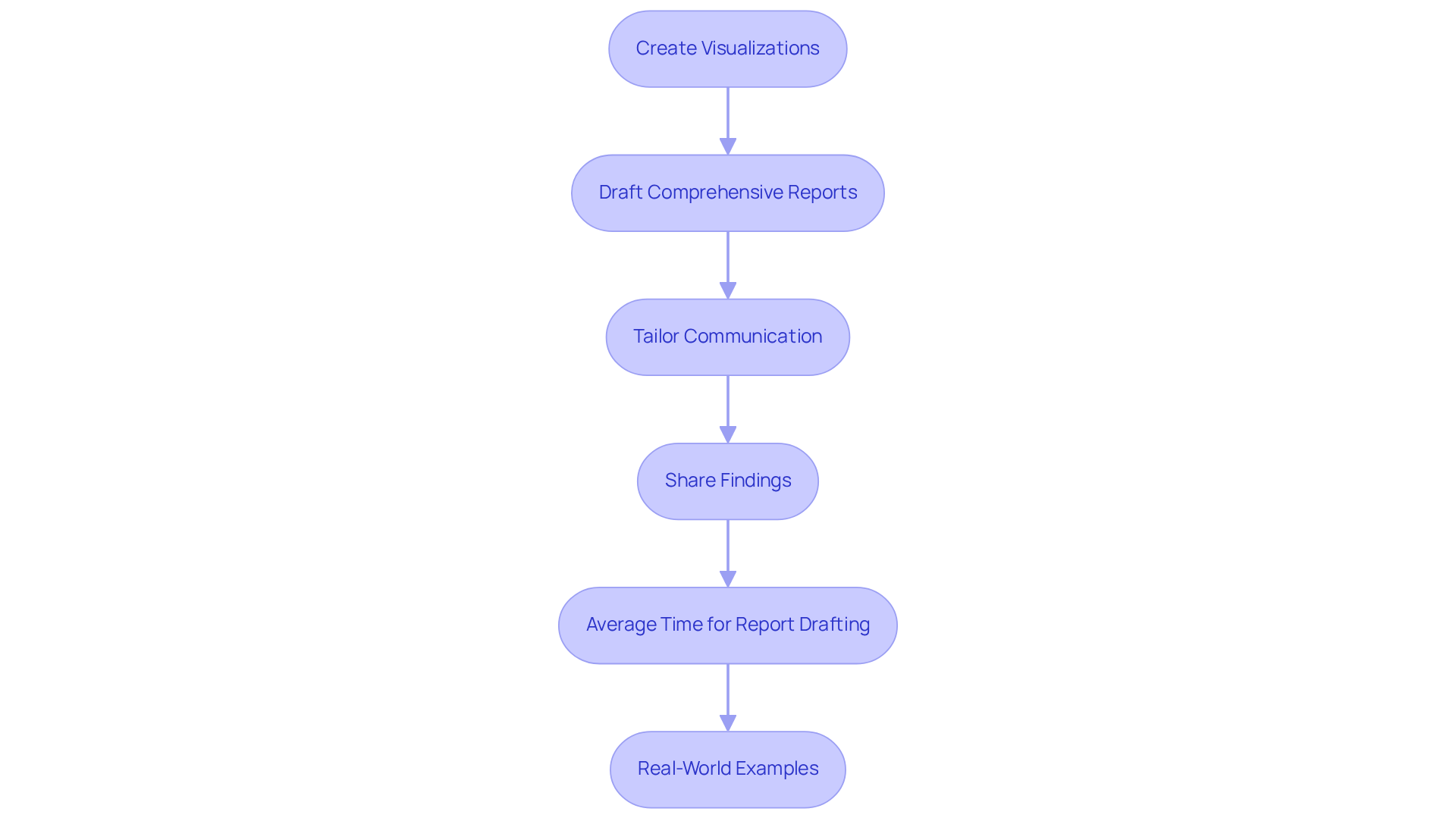
- Create Visualizations: Utilize tools such as R or Python to generate informative visualizations, including heatmaps and scatter plots, that emphasize key findings from your analysis. Visual tools enhance the interpretability of complex information.
- Draft Comprehensive Reports: Create thorough reports that summarize your methods, findings, and interpretations. Ensure that these reports include visualizations to substantiate your conclusions, making the data more accessible to various stakeholders.
- Tailor Communication: Adapt your reporting style to suit your audience. It is crucial to explain technical details clearly, especially for non-expert stakeholders, to enhance understanding of the findings.
- Share Findings: Present your findings in meetings or through publications, highlighting the implications of your results for clinical practice or further research. Effective communication can drive the application of your findings in real-world contexts.
- Average Time for Report Drafting: On average, preparing DNA analysis reports takes roughly 7.3 hours per case, as indicated by participating laboratories in the Medical Genome Initiative. This time commitment underscores the need for streamlined processes in DNA reporting.
- Real-World Examples: Implementing structured reporting practices has been shown to improve physician understanding of genetic profiling results, thereby enhancing the impact of genetic tests on patient care. For example, clear communication between ordering doctors and genetic data reporters is essential to minimize uncertainties and enhance the use of genetic tests in clinical settings.
By adhering to these steps, you will ensure that your findings from the genomic data analysis are communicated effectively, fostering understanding and application in practical scenarios. Ultimately, the clarity of your communication can transform genomic insights into actionable strategies for patient care.
Abschluss
Mastering genomic data analysis requires a structured approach that addresses the complexities inherent in the field. This guide highlights the importance of understanding genomic data types, sequencing technologies, and bioinformatics tools, which are essential for successful analysis. By following a clear process, individuals can better navigate the complexities of genomic data and make meaningful contributions to personalized medicine.
Throughout the article, key steps were outlined to ensure a comprehensive understanding of genomic data analysis. These include:
- Collecting relevant data
- Performing quality checks
- Processing data for analysis
- Conducting exploratory data analysis
- Effectively visualizing and reporting findings
Each step is designed to enhance the accuracy and reliability of the analysis, ultimately leading to more informed decisions in genomic research.
The significance of genomic data analysis cannot be overstated, as it plays a crucial role in transforming healthcare through personalized medicine. By adhering to ethical standards and utilizing advanced tools, researchers can ensure the integrity of their findings. This process not only deepens understanding of genetic information but also sparks innovation in clinical practice. By embracing these structured steps, individuals can significantly enhance their contributions to genomics, ultimately leading to improved patient outcomes and advancements in scientific knowledge.
Häufig gestellte Fragen
Why is understanding genomic data important?
Understanding genomic data is crucial for effectively interpreting sequencing results and their implications for health.
What are the main types of genomic data?
The main types of genomic data include DNA sequences, RNA sequences, and variant call formats (VCF).
What sequencing technologies should one be familiar with?
One should be familiar with sequencing methods such as Sanger sequencing and Next-Generation Sequencing (NGS), with NGS being particularly transformative for genetic studies due to its high-throughput capabilities.
What role do bioinformatics tools play in genomic data analysis?
Proficiency in bioinformatics tools, including software and coding languages like R and Python, is essential for evaluating and interpreting complex genetic information.
What is the expected trend regarding healthcare workers and bioinformatics tools by 2026?
By 2026, it is expected that 70% of healthcare workers will utilize bioinformatics tools in genetic studies, indicating the growing role of technology in healthcare.
How does TrakGene contribute to genomic data management?
TrakGene enhances the integration of genomic data by providing comprehensive electronic health records related to genetics, including cancer risk tools, which streamline the management of genetic information.
What is the first step in effective genomic data analysis?
The first step is to identify information sources, such as public databases like NCBI and Ensembl, or institutional repositories that hold valuable genetic collections.
How should datasets be selected for genomic analysis?
Datasets should be selected based on their relevance to the research question or clinical needs, ensuring they are comprehensive and ethically sourced.
Why is compliance with ethical guidelines important in genomic data collection?
Compliance with ethical guidelines and regulations, such as informed consent and privacy laws (e.g., HIPAA and GDPR), is critical for maintaining trust and integrity in genomic research.
What steps should be taken after collecting genomic data?
After collecting genomic data, it should be downloaded in suitable formats (e.g., FASTQ, BAM) and organized methodically to facilitate straightforward access during examination.
List of Sources
- Understand the Fundamentals of Genomic Data Analysis
- Scientists sharpen genetic maps to help pinpoint DNA changes that influence human health traits and disease risk (https://jax.org/news-and-insights/2026/february/scientists-sharpen-genetic-maps-to-help-pinpoint-dna-changes-that-influence-human-health-traits-and-disease-risk)
- TCGA whole-genome insights to transform oncology – SelectScience news (https://selectscience.net/article/inocras-and-broad-institute-researchers-will-release-novel-insights-from-tcga-cancer-whole-genome)
- Why genomic healthcare data matters in the development of new therapies – Drug Discovery World (DDW) (https://ddw-online.com/why-genomic-healthcare-data-matters-in-the-development-of-new-therapies-20235-202211)
- Genomic Data Analysis and Interpretation Market Report 2026 (https://researchandmarkets.com/reports/5983726/genomic-data-analysis-interpretation-market-report?srsltid=AfmBOopYYwuK6CZJ6X_9uNHLGEMKtIcHr0vuUCdzFp1uDSiQVYMRk8E5)
- Genomes in clinical care – npj Genomic Medicine (https://nature.com/articles/s41525-024-00402-2)
- Collect Relevant Genomic Data
- CDC Traveler-Based Genomic Surveillance Program Tops One Million Participants, Giving Clinical Labs Earlier Warning on Emerging Variants (https://darkdaily.com/2026/02/09/cdc-traveler-based-genomic-surveillance-program-tops-one-million-participants-giving-clinical-labs-earlier-warning-on-emerging-variants)
- Several States Introduce New Genetic Privacy Bills in Early 2026 (https://insideprivacy.com/health-privacy/several-states-introduce-new-genetic-privacy-bills-in-early-2026)
- NIH Seeks Input on Draft Policies to Protect Human Participant Research Data (https://faseb.org/journals-and-news/washington-update/nih-seeks-input-on-draft-policies-to-protect-human-participant-research-data)
- WHO releases new principles for ethical human genomic data collection and sharing (https://who.int/news/item/20-11-2024-who-releases-new-principles-for-ethical-human-genomic-data-collection-and-sharing)
- Perform Data Quality Checks and Cleaning
- NeoGenomics to Spotlight AI-Driven Genomic–Clinical Data Integration at AACR 2026 (https://clpmag.com/disease-states/cancer/neogenomics-spotlight-ai-driven-genomic-clinical-data-integration-aacr-2026)
- How the All of Us Genomic data are organized (https://support.researchallofus.org/hc/en-us/articles/29475228181908-How-the-All-of-Us-Genomic-data-are-organized)
- Practical impacts of genomic data “cleaning” on biological discovery using surrogate variable analysis – PubMed (https://pubmed.ncbi.nlm.nih.gov/26545828)
- NGS Data Quality Control: Best Practices for Accuracy – Euformatics (https://euformatics.com/blog-post/ngs-data-quality-control-best-practices-for-accuracy)
- Process Genomic Data for Analysis
- Bioinformatics in 2026: Genomics & AI-Driven Insights (https://nextmsc.com/blogs/bioinformatics-in-2026-from-genomics-to-real-world-impact)
- Improved tumor-only variant calling and mutation burden estimation with VarNet-T – Nature Communications (https://nature.com/articles/s41467-026-71705-4)
- Benchmarking SNP-Calling Accuracy Against Known Citrus Pedigrees Reveals Pangenome Advantages Over Linear References (https://biorxiv.org/content/10.64898/2026.04.07.716967v1.full)
- New tool for examining cancer genomic data could improve treatment – ecancer (https://ecancer.org/en/news/27768-new-tool-for-examining-cancer-genomic-data-could-improve-treatment)
- AI Transforms Genomic Data Analysis in 2026 | Neil Ward posted on the topic | LinkedIn (https://linkedin.com/posts/neilward_expert-predictions-for-healthtech-in-2026-activity-7402068440750125056-IQ5i)
- Conduct Exploratory Data Analysis and Modeling
- Illumina launches powerful software for connected, intuitive, and scalable multiomic analysis (https://investor.illumina.com/news/press-release-details/2026/Illumina-launches-powerful-software-for-connected-intuitive-and-scalable-multiomic-analysis/default.aspx)
- New review on statistical and machine learning tools to advance… (https://genomicsengland.co.uk/news/new-review-on-statistical-and-machine-learning-tools-to-advance-equity-in-genomic-research)
- New method to analyze complex genetic data could be the key to tackling rare diseases (https://sciencedaily.com/releases/2024/10/241030150030.htm)
- New statistical method improves genomic analyses | Penn State University (https://psu.edu/news/eberly-college-science/story/new-statistical-method-improves-genomic-analyses)
- Visualize and Report Genomic Analysis Findings
- ESMO Recommendations on clinical reporting of genomic test results for solid cancers (https://sciencedirect.com/science/article/pii/S0923753424010111)
- Best practices for the interpretation and reporting of clinical whole genome sequencing – npj Genomic Medicine (https://nature.com/articles/s41525-022-00295-z)
- Biological Data Visualization Market Size, Trends & Growth Report by 2034 (https://straitsresearch.com/report/biological-data-visualization-market)
- Genomic Data Analysis and Interpretation Market Report 2026 (https://researchandmarkets.com/reports/5983726/genomic-data-analysis-interpretation-market-report?srsltid=AfmBOopm47lNlUzntQEdRPkJApKGwQYnaEGTjhmRJ_deOFbH4gKFGghL)






